Q&A: What is biophysics?
Biomolecules such as proteins and DNAs have been selected by evolution to perform specific biological functions. The molecular architectures and the biological processes are constrained by the laws of physics and chemistry. For example, the folded structures of proteins and the DNA double helix are held together by physical forces. Proteins interact with each other and with DNAs by electrostatic and other types of forces. The interaction forces can be modulated by changing protein and DNA sequences (i.e., mutations).
FSU has a rich biophysics research environment, both in the Physics Department and in the Institute of Molecular Biophysics (IMB). The Graduate Program in Molecular Biophysics (MOB), which is housed in IMB, incorporates a multidisciplinary research focus. (Note that application deadlines for the Molecular Biophysics Program may differ from Physics.) The biophysics faculty within the Department of Physics has close interactions with faculties in the Department of Chemistry & Biochemistry, Department of Biological Science, the College of Medicine, and the Institute of Molecular Biophysics.
Click a menu option to jump or scroll down for more information on biophysical research by Physics faculty
- Research Overview
- Protein-protein interactions
- Protein folding
- Protein stability
- Biomotors
- Hybrid nanostructures and devices
- Dip-Pen Nanolithography and Surface Diffusion
- Gel Formation
- Faculty
Research Overview
Prof. David van Winkle's lab uses a wide range of experimental techniques to study biophysical problems in crystals, gels, and liquid crystals. In addition, Prof. Bernd Berg and Prof. Per Arne Rikvold, with backgrounds in high-energy and condensed-matter physics, are branching into biophysics and are using their expertise in developing Monte-Carlo algorithms and other statistical mechanics methods to study biological processes such as protein folding.
There are many opportunities for graduate and undergraduate research in biophysics at FSU. Prof. Hong's lab uses atomic force microscopy to manipulate biomolecules into functional nano-structures. Prof. Zhou's lab uses computer modeling to study protein folding, stability, and protein-protein and protein-DNA interactions. Prof. van Winkle's lab uses a wide range of experimental techniques to study biophysical problems in crystals, gels, and liquid crystals. In addition, Prof. Berg and Prof. Rikvold, with backgrounds in high-energy and condensed- matter physics, are branching into biophysics and are using their expertise in developing Monte-Carlo algorithms and other statistical mechanics methods to study biological processes such as protein folding.
FSU has a rich biophysics research environment. The biophysics faculty within the Department of Physics has close interactions with faculties in the Departments of Chemistry and Biochemistry and Biological Science, the School of Medicine, and the National High Magnetic Field Laboratory (NHMFL). These interations are forstered by several interdisciplenary organizations, such as the Institute of Molecular Biophysics, the School of Computational Science and Information Technology (CSIT), and the Center for Materials Research and Technology (MARTECH).
Protein-protein interactions
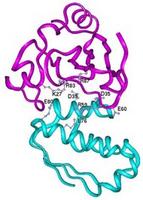 Biophysics Proteins |
Interactions between proteins are essential for biological functions. Our research aims to gain fundamental understanding of the determinants of binding affinity and binding rates. Based on statistical mechanics, we develop physical models that can be conveniently implemented on the computer to realistically predict binding properties. An example is the transition-state theory we developed to predict the enhancement of protein association rates by electrostatic interactions. Other questions being addressed include:
|
Protein Folding
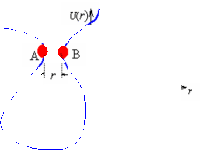 Biophysics Folding |
The folding of a protein molecule can be viewed as the accumulation of native contacts. Based on modeling the unfolded protein molecule as a polymer chain, we have developed a theory for the rate of forming native contacts. This theory is now being extended and applied to study folding processes such as beta-hairpin and coiled-coil formation. |
Protein Stability
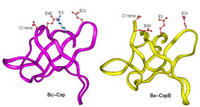 Biophysics Protein Stability |
The major stabilizing forces of protein structures are hydrophobic and electrostatic. While there is consensus on the hydrophobic contributions, the roles of electrostatic interactions in protein stability have been uncertain. We have made significant progress in modeling electrostatic effects. We are now developing better solvation models using molecular dynamics simulations with explicit solvent and investigating effects of charges and electrostatic interactions in protein folding, aggregation, and amyloid formation. |
Biomotors
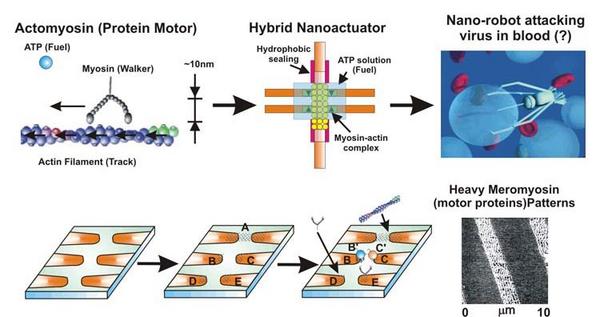 Biophysics Biomotors |
|
Biomotors are protein molecules that convert chemical energy to mechanical motion. They generate basic motions for many moving biological systems including our muscles. One example is actomyosin that is comprised two components: actin and myosin. Actin provides a track, on which myosin walks while consuming chemical energy from ATP hydrolysis. The biomotor is a fascinating masterpiece of the Nature. First of all, each motor unit has a size of ~ 10 nm, which is a thousand times smaller than any man-made motors. Moreover, the fuel efficiency of biomotors is higher than any existing man-made engines. An interdisciplinary research team at Florida State University is performing extensive research to understand biomotors. Furthermore, utilizing newly developed nanotechnology, new hybrid engines based on biomotors are being developed. This work is supported by DARPA. |
Hybrid Nanostructures and Devices
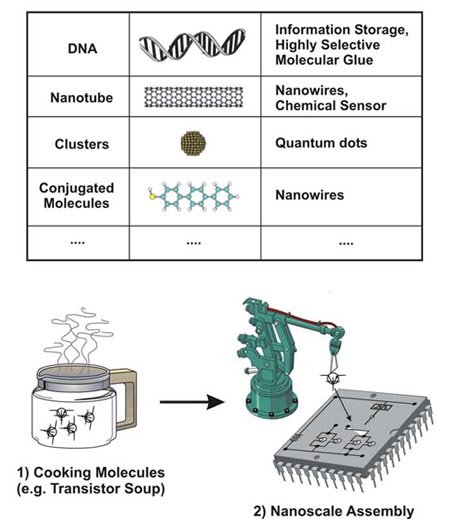 Biophysics Nanostructures |
| Due to dramatic progress of nanotechnology and biological science, biological molecules (e.g. DNA and proteins) can now be combined with solid nanostructures (e.g. nanoparticles and nanowires) to build a new generation of devices. These include biological sensors and nanomechanical systems based on protein motors. These new types of devices are called hybrid devices. An interdisciplinary research team at FSU is trying to understand the properties of hybrid nanostructures and to develop various hybrid devices. |
Dip-Pen Nanolithography and Surface Diffusion
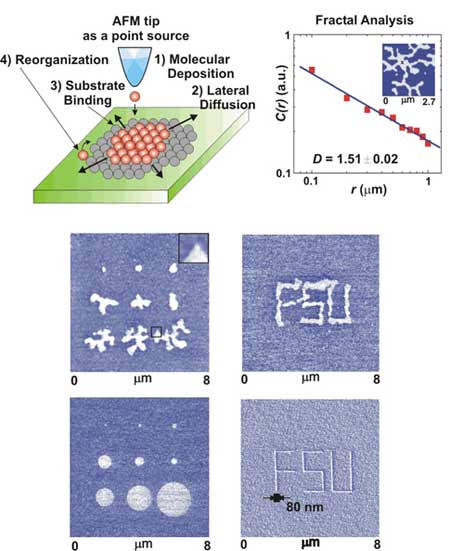 Biophysics Lithography |
|
Surface Diffusion and Nanopatterning The diffusion behavior of organic molecules on solid surface is both a very intriguing statistical physics problem and very important in practical applications. FSU researchers have a tool named "dip-pen nanolithography" which allows the deposition of organic molecules in a nanometer scale region on the solid substrate using nanometer scale dip-pen (top). When deposited onto solid substrates, organic molecules diffuse out in different ways, forming fractal-like (middle) or circular patterns (bottom). The diffusion behavior dramatically affects the nanopatterning results (See letters of "FSU"). |
Gel Formation
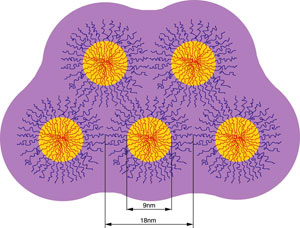 Biophysics Gels |
The accompanying picture is a representation of the close packed structure formed by micelles of a tri-block copolymer. The material (F-127 from BASF) consists of a poly-propyleneoxide (PPO) core with poly-ethyleneoxide (PEO) chains attached to both ends of the PPO. At room temperature the PPO is relatively hydrophobic compared with the PEO. In aqueous solution, PPO cores from many molecules aggregate. The molecules form micelles approximately 18 nm in diameter. At high enough concentration, the micelles pack into a hexagonal lattice and the material is a non-crosslinked gel. We discovered that electrophoretic separations on such gels are possible, and that a whole host of new separations phenomena arise. As an aside, when the temperature is lowered below 17 degrees C, the PPO and PEO exhibit about the same relative hydrophobic character. The micelles disassemble and the solution becomes a free-flowing liquid. Thus it appears to melt on cooling and freeze on warming. |
Find out more about Biophysics at FSU at:
Institute of Molecular Biophysics (IMB)
